Homework 7
Up to the Phylogenetics main page
Conditioning on variability in the Mk model
The python script below will be modified by you to compute the site likelihoods of a 4-taxon tree for 2 characters using the Mk 2-state model. The first of the two characters is constant and the second character is variable. You will use the site likelihood for the constant character to condition the site likelihood for the variable character on the fact that it is variable. You will check your results using PAUP*.
Be sure to ask me for help if you get stuck.
What to do
Login to your account on the Storrs HPC cluster and create a file named hw7.py using nano containing the code below.
from math import exp, log
# You do *not* need to modify this function
# same_list holds the number of edges with the same state at both ends
# over all 4 possible combinations of ancestral states
# v is the edge length (same for all 5 edges in the tree)
def sitelike(same_list, v):
probsame = 0.5 + 0.5*exp(-2.*v)
probdiff = 0.5 - 0.5*exp(-2.*v)
like = 0.0
for n in same_list:
nsame=float(n)
ndiff=float(5-n)
like += 0.5*(probsame**nsame)*(probdiff**ndiff)
return like
# 00 01 10 11 <-- ancestral state combinations possible
same1 = [ 5, 2, 2, 1] # no. edges with same state at both ends
same2 = [ 3, 4, 0, 3] # in tree 1 (same1) and tree 2 (same2)
# Compute the site likelihood for each of the two characters
# by calling the function sitelike (we'll assume v = 0.1 for
# all edges.
v = 0.1
like1 = sitelike(same1, v)
like2 = sitelike(same2, v)
print('Mk model (edge lengths all %.1f):' % v)
print(' likelihood (char. 1) = %.5f' % like1)
print(' likelihood (char. 2) = %.5f' % like2)
# TODO: calculate likelihood for character 2 conditioned on variability
#like2cv = xxxx
#print('Mk model (edge lengths all %.1f, conditioning on variability):' % v)
#print(' likelihood (char. 2) = %.5f' % like2cv)
#print(' log-likelihood (char. 2) = %.5f' % log(like2cv))
Ensure that you have access to python3 on the cluster by loading the python module:
module load python
Run the hw7.py script as follows:
python3 hw7.py
Because the parts that are not yet completed by you are commented out (using hashtags at the beginning of the line), the hw7.py program should run and spit out the likelihood without conditioning on variability.
After running it once to make sure everything works, edit the hw7.py script in nano, uncommenting the final 4 lines and replacing the xxxx placeholder with the correct formula. Assume the data for characters 1 and 2 are as shown in Figure 1.
The sitelike function
Near the top of your hw7.py file is the definition of a function named sitelike. The function sitelike takes 2 arguments.
The first argument,same_list, is a list of 4 numbers representing the number
of edges (of the 5 total) in which the state is the same
at both ends. There are 4 such values because there are 4
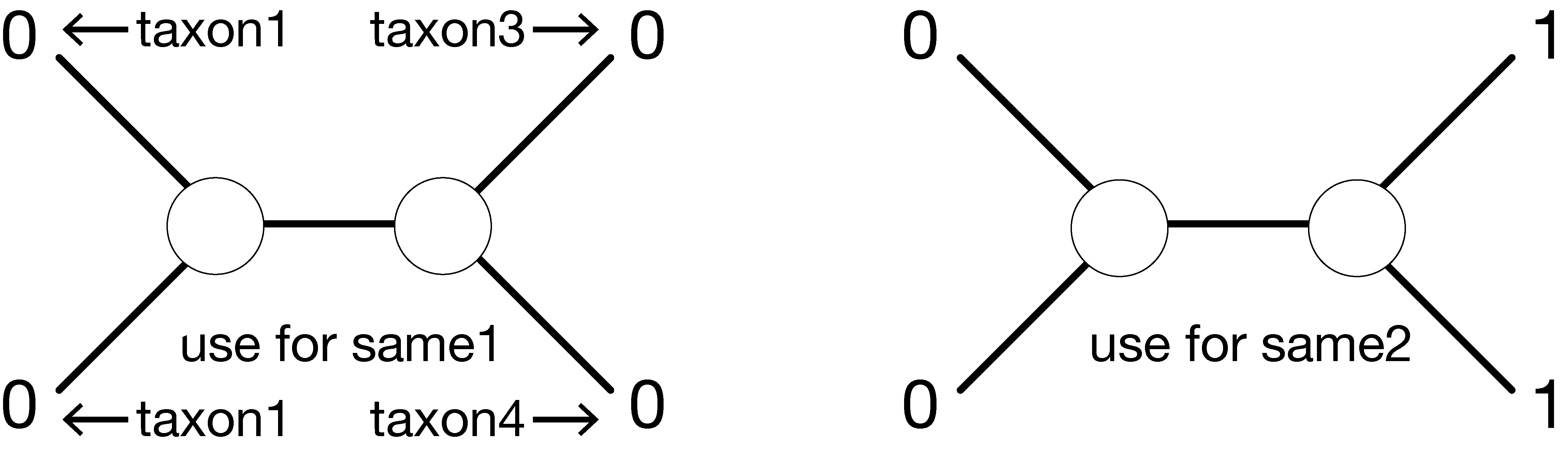
possible ancestral state combinations (00, 01, 10, 11) for 2 states (0 and 1). For example, in the tree on the left (labeled “Use for same 1”, the ancestral state combination 10 assigns 1 to the left ancestral state and 0 to the right ancestral state. This assignment results in the rightmost 2 edges (i.e. the edges connected to taxon 3 and taxon 4) having the same state at each end, and thus the array same1 has a 2 in the 3rd position corresponding to the 10 ancestral state combination.
The second argument is v, which is the edge length to be used for all 5 edges.
Checking your answer
Check your answer using PAUP*. You can get the data for this homework into PAUP* by creating a nexus file named hw7.nex in nano that has this content:
#nexus
begin data;
dimensions ntax=4 nchar=2;
format datatype=standard missing=? gap=-;
matrix
taxon1 00
taxon2 00
taxon3 01
taxon4 01
;
end;
begin trees;
tree it = [&U] (taxon1:.1,taxon2:.1,(taxon3:.1,taxon4:.1):.1);
end;
begin paup;
set crit=like;
[!
***************************************
*** Not conditioning on variability ***
***************************************
]
lset nst=1 genfreq=equal condvar=no;
lscores 1 / sitelike userbrlen;
[!
***********************************
*** Conditioning on variability ***
***********************************
]
exclude 1;
lset nst=1 genfreq=equal condvar=yes mkStateSpace=fixed;
lscores 1 / sitelike userbrlen;
quit;
end;
The paup block above has two sections. The first section computes the likelihood for each character without conditioning on variability. (Note that PAUP* requires you to use genfreq rather than basefreq if the datatype is standard.) The exclude 1 statement in the second part is needed in order to exclude the first character (because constant characters are not allowed if condvar is set to yes).
What if my answer differs from PAUP*’s answer?
First, make sure you are getting the correct site likelihoods without conditioning on variability.
Second, if you are getting the correct site likelihoods when not conditioning on variability but not matching when you do condition on variability, be sure you are correctly calculating the probability of the site being variable. This value is 1 minus the probability that a site is constant, and remember that there are two ways that a site can be constant (either all 0s or all 1s at the leaves). You have calculated the probability that a site has all 0s (this is stored in the like1 variable), and the probability of a site having 1 at every leaf is the same, so all you need to do is multiply the value stored in like1 by 2 to get the probability that a site is constant.
What do I turn in?
Please send me a direct message in Slack and attach both your python program hw7.py as well as your hw7.nex file.